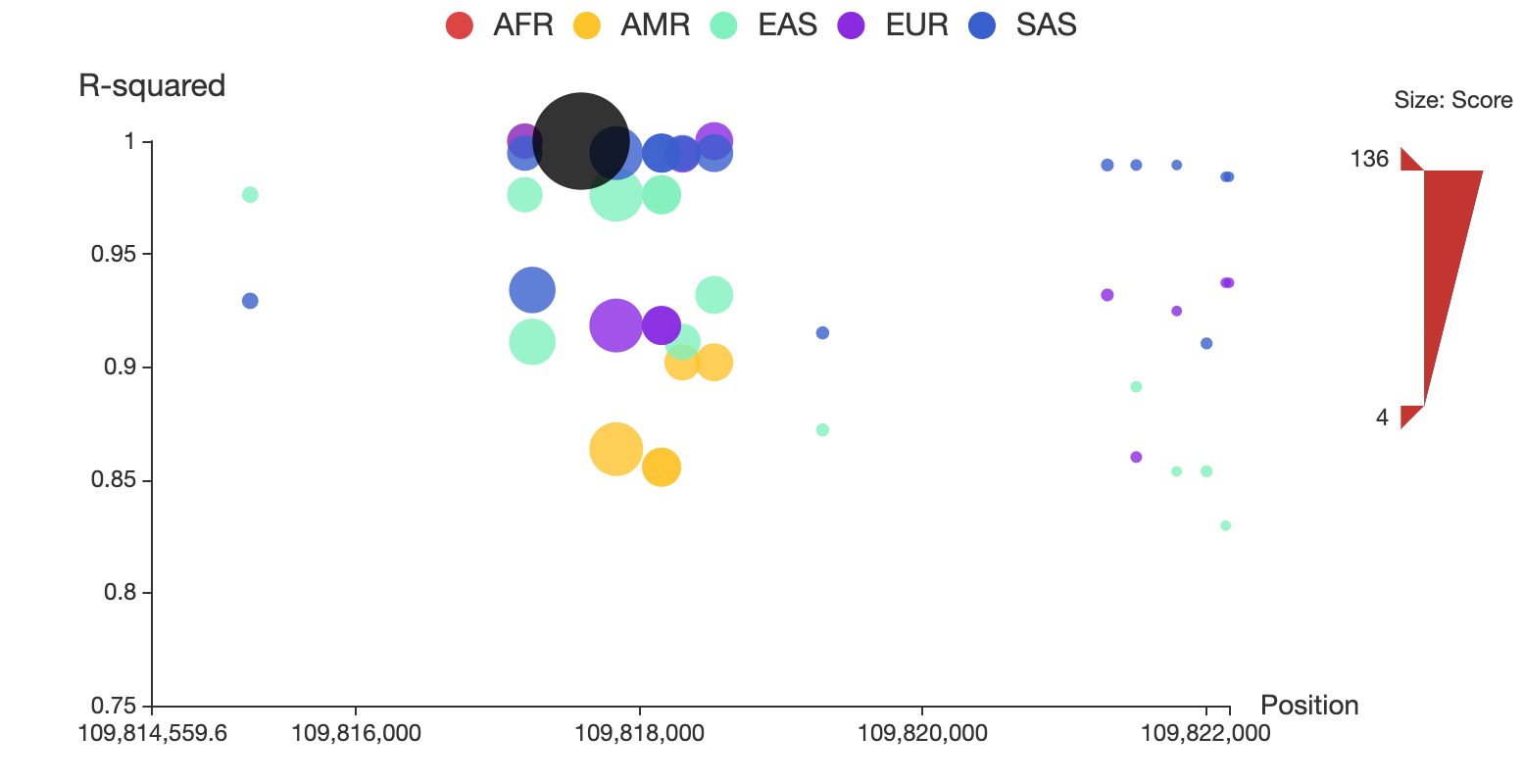
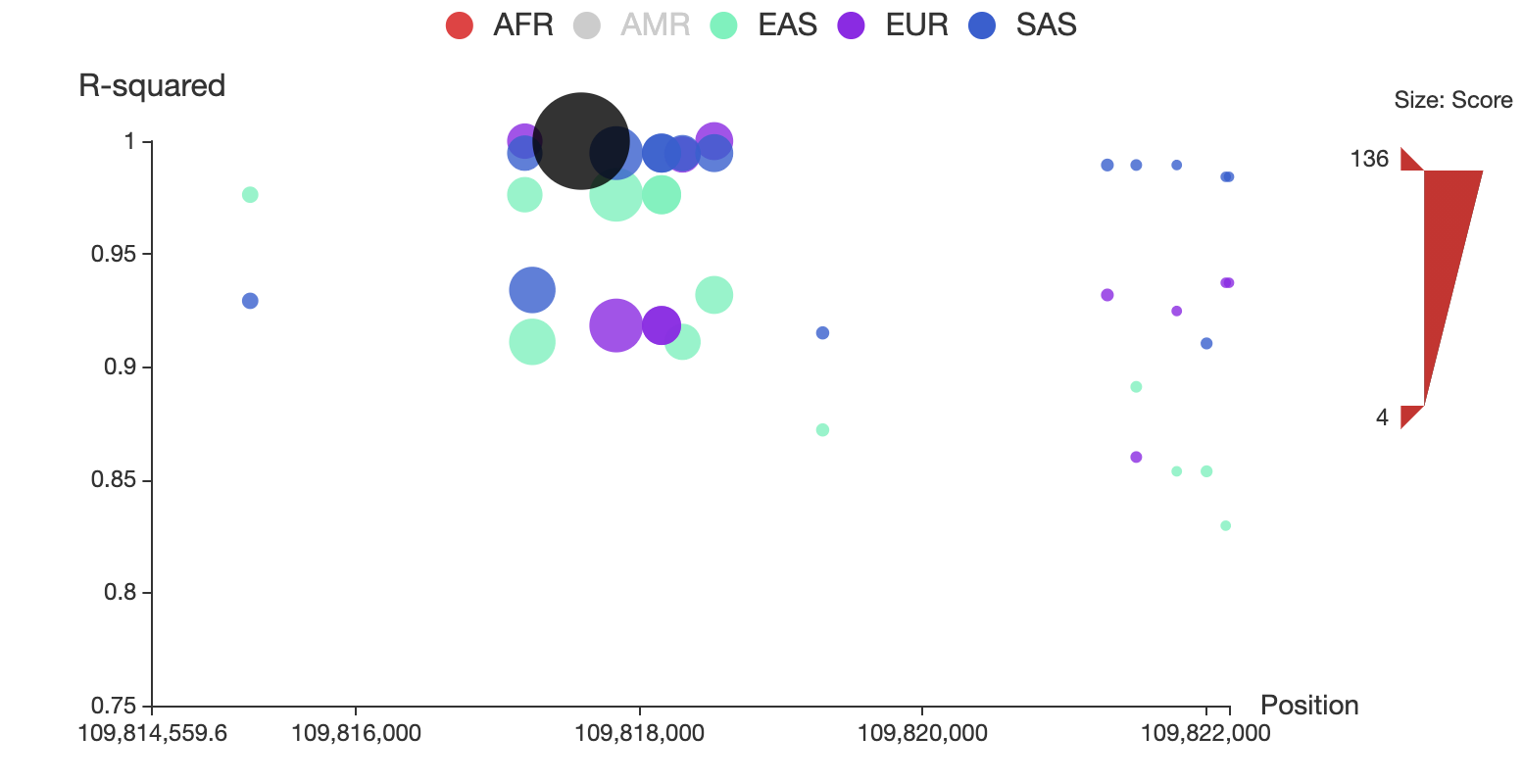
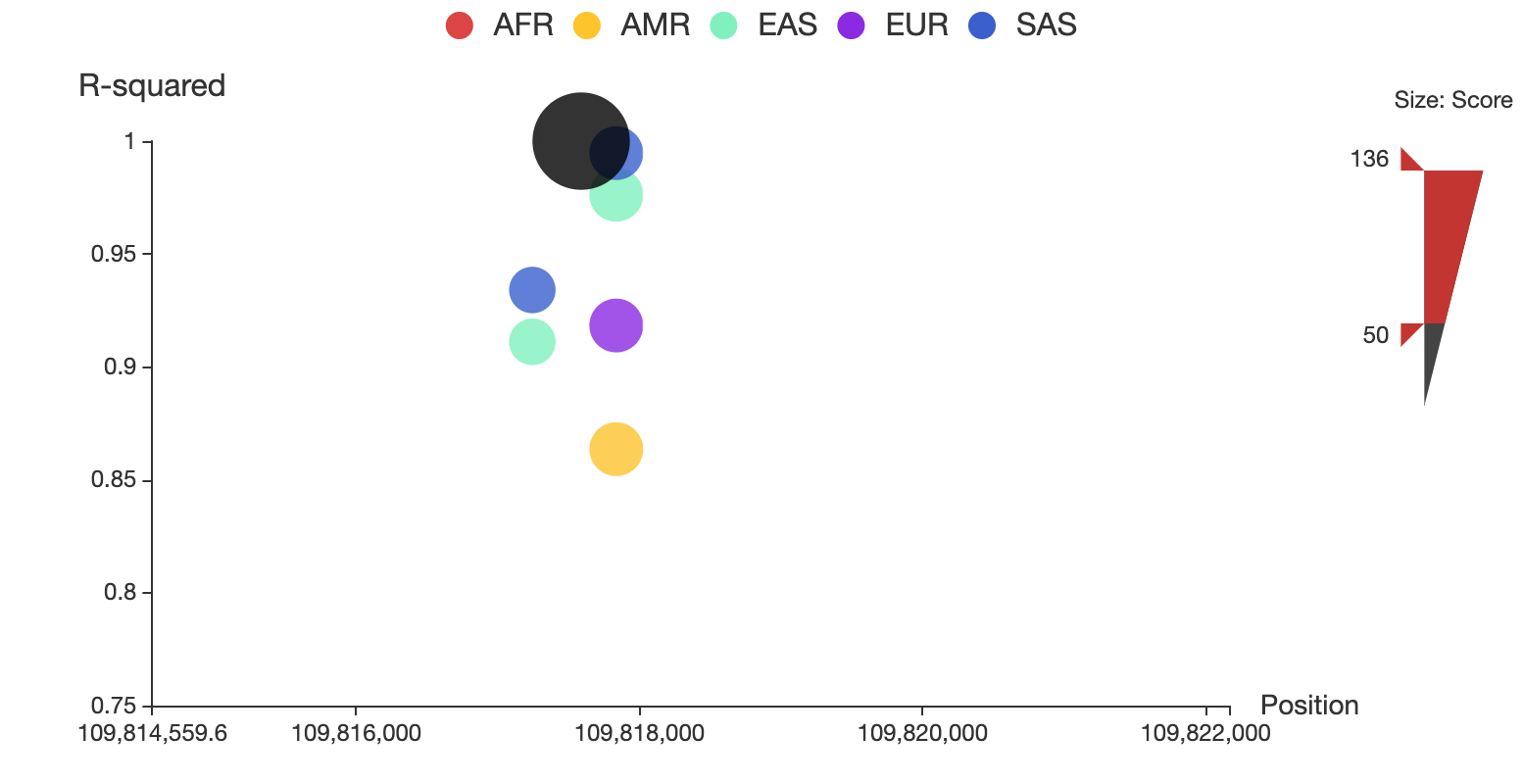
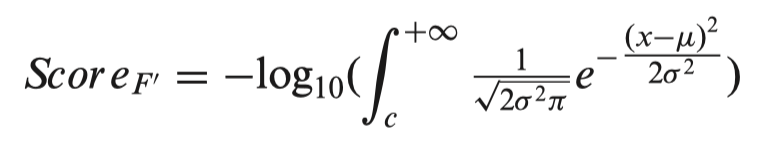3dsnp v2 for developers
3DSNP v2 extends the API functions of the previous version. The domain was changed and more importantly, SV data and some new tables were included. Now data can be accessed by three means: SNP ids, SV ids or Chromatin position.
We recommend using positions to search for variants which will display both SNPs and SVs in the target region.
We marked new features with *
Note: The original API are always open.
Overview
URL
https://omic.tech/3dsnpv2/api.do
Format supported
JSON/XML
HTTP request method
GET/POST
Login required
No
Data access restrictions
Frequency limit: No
Request
Request parameters
|
Required |
Type |
Information |
| id*/position |
true |
string |
Represents the SNP/SV ID or genomic position, at least one of them is required,
multiple SNP IDs or positions should separated by comma ‘,’.
Dash symbol ‘-‘
The format of parameter ‘position’
should be ‘1000000-1000100’. SV IDs could be found in HGSVC v2. |
| chrom |
false |
string |
Represents the chromosome of queried position and is required when parameter ‘position’ is used.
When there are more than one positions, the corresponding chromosomes should also be separated by ‘,’. |
| type |
true |
string |
Data type for searching, multiple types should be separated by comma ‘,’.
Available types are listed below. |
| format |
true |
string |
Represents data types returns. Json and XML formats are supported. |
Request data type
| DataType |
Description |
| basic |
Basic information of SNP, including sequential facts and phenotype from 1000G project. |
| chromhmm |
Chromatin state information generated by the core 15-state ChromHMM models trained
across a variety of cell types. |
| motif |
Transcription factor binding motifs altered by SNP. |
| tfbs |
Transcription factor binding sites in a variety of cell types. |
| eqtl |
Expression quantitative trait loci (eQTL). |
| 3dgene |
Genes that interact the query SNP through chromatin loops. |
| 3dsnp |
SNPs that interact the query SNP through chromatin loops. Not available for the query of position. |
| phylop |
PhyloP scores of genomic region surrounding the query SNP. |
| ccre* |
The status of open chromatin for over 750,000 candidate cis-regulatory elements (cCREs) in 54 distinct cell types. |
| genetics* |
Integrated haplotype scores (iHS) and Fixation index (Fst) for five continental population obtained from 1000 Genomes Phase 3 (final phase) |
| clinvar* |
ClinVar aggregates information about genomic variation and its relationship to human health. |
Response
Response parameters
|
Type |
DataType |
Description |
| id |
string |
basic |
SNP ID |
| chr |
string |
basic |
Chromosome name |
| position |
string |
basic |
Location of the query |
| MAF |
string |
basic |
Minor allele frequency |
| Ref |
string |
basic |
Reference Allele |
| Alt |
string |
basic |
Alternative Allele |
| EAS |
string |
basic |
Allele frequency in the EAS populations |
| AMR |
string |
basic |
Allele frequency in the AMR populations |
| AFR |
string |
basic |
Allele frequency in the AFR populations |
| EUR |
string |
basic |
Allele frequency in the EUR populations |
| SAS |
string |
basic |
Allele frequency in the SAS populations |
| linearClosestGene |
string |
basic |
Linear cloest genes |
| data_gene |
JsonArray |
basic |
listed below in JsonArray Parameters |
| chromhmm |
string |
chromhmm |
Chromatin state from ChromHMM core 15-state model |
| data_chromhmm |
JsonArray |
chromhmm |
listed below in JsonArray Parameters |
| motif |
string |
motif |
Sequence motif altered by the query SNP |
| data_motif |
JsonArray |
motif |
listed below in JsonArray Parameters |
| tfbs |
string |
tfbs |
Transcription factor binding sites the query locates |
| data_tfbs |
JsonArray |
tfbs |
listed below in JsonArray Parameters |
| eqtl |
string |
eqtl |
Expression quantitative trait loci |
| data_eqtl |
JsonArray |
eqtl |
listed below in JsonArray Parameters |
| data_loop_gene |
JsonArray |
3dgene |
listed below in JsonArray Parameters |
| data_loop_snp |
JsonArray |
3dsnp |
listed below in JsonArray Parameters |
| physcores |
string |
physcores |
PhyloP scores of the query SNP and its +/-10 bp adjacent regions |
| ccre.position* |
string |
ccre |
The corresponding peak position of cCREs |
| mapping* |
string |
ccre |
Mapping rate of cCREs |
| Fol/Acn/Skm1/…/Swn_2* |
string |
ccre |
cCREs in 54 distinct cell types |
| Fst_EUR* |
string |
genetics |
Fixation index in EUR |
| Fst_SAS* |
string |
genetics |
Fixation index in SAS |
| Fst_EAS* |
string |
genetics |
Fixation index in EAS |
| Fst_AMR* |
string |
genetics |
Fixation index in AMR |
| Fst_AFR* |
string |
genetics |
Fixation index in AFR |
| iHS_EUR* |
string |
genetics |
Integrate Haplotype score in EUR |
| iHS_SAS* |
string |
genetics |
Integrate Haplotype score in SAS |
| iHS_EAS* |
string |
genetics |
Integrate Haplotype score in EAS |
| iHS_AMR* |
string |
genetics |
Integrate Haplotype score in AMR |
| iHS_AFR* |
string |
genetics |
Integrate Haplotype score in AFR |
| xpnsl_EUR* |
string |
genetics |
cross-population NSL in EUR |
| xpnsl_SAS* |
string |
genetics |
cross-population NSL in SAS |
| xpnsl_EAS* |
string |
genetics |
cross-population NSL in EAS |
| xpnsl_AMR* |
string |
genetics |
cross-population NSL in AMR |
| xpnsl_AFR* |
string |
genetics |
cross-population NSL in AFR |
| ClinVarID* |
string |
clinvar |
the ClinVar Allele ID |
| CLNDN* |
string |
clinvar |
ClinVar’s preferred disease name for the concept specified by disease identifiers in CLNDISDB |
| CLNDISDB* |
string |
clinvar |
Tag-value pairs of disease database name and identifier |
| CLNREVSTAT* |
string |
clinvar |
ClinVar review status for the Variation ID |
| CLNSIG* |
string |
clinvar |
Clinical significance for this single variant |
| CLNSIGCONF* |
string |
clinvar |
Conflicting clinical significance for this single variant |
| CLNVC* |
string |
clinvar |
Variant type |
| CLNVCSO* |
string |
clinvar |
Sequence Ontology id for variant type |
| CLNVI* |
string |
clinvar |
the variant’s clinical sources reported as tag-value pairs of database and variant identifier |
| GENEINFO* |
string |
clinvar |
Gene(s) for the variant reported as gene symbol:gene id |
| MC* |
string |
clinvar |
comma separated list of molecular consequence in the form of Sequence Ontology ID|molecular_consequence |
| ORIGIN* |
string |
clinvar |
Allele origin |
JsonArray Parameters
|
Type |
JsonArray |
Description |
| geneID |
string |
data_gene |
RefSeq Gene ID |
| geneName |
string |
data_gene |
Official gene symbol |
| geneRelativePosition |
string |
data_gene |
Relative position of the closest gene to the query |
| geneDescription |
string |
data_gene |
Gene description |
| chromhmmCell |
string |
data_chromhmm |
Cell type of the corresponding chromatin state |
| chromhmmName |
string |
data_chromhmm |
Short name of chromatin state |
| chromhmmFullName |
string |
data_chromhmm |
Full name of chromatin state |
| chromhmmCellDescription |
string |
data_chromhmm |
Cell type description |
| chromhmmTissue |
string |
data_chromhmm |
Tissue of the cell type |
| motif |
string |
data_motif |
Motif ID in TRANSFAC or JASPAR |
| motifStrand |
string |
data_motif |
Strand of the motif |
| motifSource |
string |
data_motif |
Database source of the motif |
| motifMatchedSequence |
string |
data_motif |
Matched sequence for the motif |
| motifMatchedSequencePos |
string |
data_motif |
Relative position of the query to the sequence |
| motifRef |
string |
data_motif |
Reference allele |
| motifAlt |
string |
data_motif |
Alternative allel |
| tfbsCell |
string |
data_tfbs |
Cell type of the corresponding TFBS |
| tfbsFactor |
string |
data_tfbs |
Name of the transcription factor |
| tfbsCellTissue |
string |
data_tfbs |
Tissue of the cell type |
| tfbsDNAAccessibility |
string |
data_tfbs |
DNA accessibility of the TFBS |
| tfbsCellDescription |
string |
data_tfbs |
Description for the cell type |
| eqtlGene |
string |
data_eqtl |
Related gene of the eQTL |
| eqtlPValue |
string |
data_eqtl |
P-value of the eQTL |
| eqtlTissue |
string |
data_eqtl |
Tissue in which the eQTL identified |
| eqtlEffect |
string |
data_eqtl |
Effect size of the eQTL |
| loopGene |
string |
data_loop_gene |
Genes interacting the query SNP through chromatin loops |
| loopGeneID |
string |
data_loop_gene |
RefSeq Gene ID |
| loopGeneDescription |
string |
data_loop_gene |
Gene description |
| loopCell |
string |
data_loop_gene/data_loop_snp |
Cell type in which the chromatin loop was identified |
| loopCellTissue |
string |
data_loop_gene/data_loop_snp |
Tissue of the cell type |
| loopCellDescription |
string |
data_loop_gene/data_loop_snp |
Cell type description |
| loopStart |
string |
data_loop_gene/data_loop_snp |
Start genomic position of the chromatin loop |
| loopEnd |
string |
data_loop_gene/data_loop_snp |
End genomic position of the chromatin loop |
| loopType |
string |
data_loop_gene/data_loop_snp |
Type of the chromatin loop: “Within Loop” or “Anchor-to-Anchor” |
| loopSNP |
string |
data_loop_snp |
SNPs interacting with the query and in the same LD block through chromatin loops |
| loopLD |
string |
data_loop_snp |
r^2 in LD |
| loopPopulation |
string |
data_loop_snp |
Continental population (AFR, AMR, ASN, EUR and SAS) |
Request with position
URL example1 : single position and single data type in json format
Request URL :
https://www.omic.tech/3dsnpv2/api.do?position=1000000-1100000&chrom=chr11&format=json&type=basic
Response format :
[
{
"id":"chr11-1009478-INS-50",
"position":"1009477",
"chrom":"chr11",
"AFR":"0",
"AMR":"0",
"Alt":"AACACGCAGCCCATGACCCCGCGCCAGGGTCTGGAGGGACGGCCCCGGGGG",
"EAS":"0",
"EUR":"0",
"Ref":"A",
"SAS":"0",
"MAF":"INS,0.000000",
"linearClosestGene":""
},
{
"id":"rs544411125",
"position":"1000017",
"chrom":"chr11",
"AFR":"0",
"AMR":"0",
"Alt":"A",
"EAS":"0",
"EUR":"0",
"Ref":"G",
"SAS":"0.001",
"MAF":"A,0.000199681",
"linearClosestGene":"AP2A2,161,intron-variant",
"data_gene":[
{
"geneID":"161",
"geneName":"AP2A2",
"geneRelativePosition":"intron-variant",
"geneDescription":"adaptor related protein complex 2 alpha 2 subunit"
}]
}
]
URL example2 : single position and mutilple data types in xml format
Request URL :
https://www.omic.tech/3dsnpv2/api.do?position=100000-1000100&chrom=chr1&format=xml&type=eqtl,motif
Response format :
<a>
<e class="object">
<chrom type="string">chr1</chrom>
<eqtl type="string"/>
<id type="string">chr1-121118-INS-113</id>
<motif type="string"/>
<position type="string">121117</position>
</e>
<e class="object">
<chrom type="string">chr1</chrom>
<id type="string">chr1-126241-DEL-38630</id>
<position type="string">126241</position>
<data_motif class="array">
<e class="object">
<motif type="string">CEBPB_02</motif>
<motifAlt type="string">DEL</motifAlt>
<motifMatchedSequence type="string">TGATTGCACCACTG</motifMatchedSequence>
<motifMatchedSequencePos type="string">16992</motifMatchedSequencePos>
<motifRef type="string">.</motifRef>
<motifSource type="string">Transfac</motifSource>
<motifStrand type="string">-</motifStrand>
</e>
<e class="object">
<motif type="string">ETS1_B</motif>
<motifAlt type="string">DEL</motifAlt>
<motifMatchedSequence type="string">GCAGGAAGTCAGGGA</motifMatchedSequence>
<motifMatchedSequencePos type="string">-27799</motifMatchedSequencePos>
<motifRef type="string">.</motifRef>
<motifSource type="string">Transfac</motifSource>
<motifStrand type="string">+</motifStrand>
</e>
</data_motif>
<eqtl type="string"/>
<motif type="string">Transfac,CEBPB_02,-,TGATTGCACCACTG,16992;Transfac,ETS1_B,+,GCAGGAAGTCAGGGA,-27799;Transfac,CEBPB_01,+,GGGTGAGGCAAGGG,-10490;Transfac,EBF_Q6,-,TTCCCTTGAGA,32414;Transfac,KROX_Q6,-,CTCGCCCCCTCCTC,4826;Transfac,CEBP_Q2_01,+,GTTGCCCAAGCT,-24111;Transfac,MTF1_Q4,-,ACTGCGCCCAGCCT,37618;Jaspar,SPI-1,-,CGGAAG,3705;Transfac,MYOD_Q6_01,-,TTGAAGCAGGTGATGGAG,24991;Transfac,TEL2_Q6,-,CCACTTCCTG,32686;Transfac,CRX_Q4,+,CCCGTAATCCCAG,-27209;Transfac,R_01,-,TGGGCCACCGGATGTGGTCCT,5445;Transfac,HNF4_01,-,ACGCGGACAGAGGTCAGCG,10966;Transfac,PAX4_01,+,GGAGGTGACCCGTGGGCAGCC,-6023;Transfac,PAX4_02,+,GAATAATTGCC,-1320;Transfac,PAX4_03,-,AGCCCCCACCCC,8402;Transfac,PAX4_04,+,AAAAATTAGCCGGGTGTGGTGGCACACACC,-3883;Transfac,IK3_01,+,TACTGGGAATGTC,-16898;Jaspar,SAP-1,-,ACCGGATGT,5439;Transfac,E2F1_Q4,+,CTTGGCGG,-33552;Transfac,HNF1_Q6,-,AGGTTAATAATTATCTCT,35228;Transfac,E2F1_Q3,+,CGTGGCGC,-28392;Transfac,AR_02,-,CGCCCACGATCAACGTGTTCTGTTCTG,8539;Transfac,ETF_Q6,+,GCGGCGG,-11412;Transfac,EN1_01,-,GTAGTGG,3310;Transfac,SREBP_Q3,-,CCCATCACCCCA,17405;Transfac,AP4_01,-,AGGATCACCTGAGGTCAG,3413;Transfac,HAND1E47_01,+,GGTGGTGTCTGGCACT,-5938;Transfac,E2F1_Q3_01,-,TGGGCGGCAGCAGGGC,6056;Transfac,STAT3_01,-,GGTGATTTCCAGGATGTGAGC,17822;Transfac,MYB_Q3,+,GGTGCCAGTTG,-7224;Transfac,HMEF2_Q6,-,GGCTAAAACTACCCCT,35670;Transfac,EGR2_01,-,TCACGTGGGCGG,6061;Transfac,E2F_Q2,-,GGCGCG,6794;Transfac,PAX8_01,-,CGGTGTCGAGTGAGG,13827;Transfac,RP58_01,-,AACACATCTGGA,37199;Transfac,CEBPGAMMA_Q6,-,CCCACTTCAGAGA,19517;Transfac,HEN1_01,+,TCGGTGCTCAGCTGAGTCTGCA,-2833;Transfac,E2_Q6_01,-,CCCACCGTCTCTGGTT,19989;Transfac,HEN1_02,-,CCTGGGCCCAGCTCCGTCCTCT,9184;Transfac,USF2_Q6,+,CACGCG,-11114;Transfac,SP1_Q6,+,CAAGGGCGGGGCC,-11202;Transfac,SMAD4_Q6,+,AGGATGCAGCCAGCT,-33630;Transfac,CIZ_01,+,GAAAAAGCC,-12404;Transfac,TAL1ALPHAE47_01,-,TTGGCCAGATGGGGTC,14330;Jaspar,deltaEF1,+,CACCTG,-3326;Transfac,POLY_C,-,GAGAAAACCCTCCTGCTG,8438;Jaspar,ARNT,+,CACGTG,-6055;Transfac,MEF3_B,-,TGCCCAGGTTTCA,28126;Transfac,GATA2_01,+,GGGGATGGGG,-6520;Transfac,GR_01,+,GCAGCATGGGCAGGATGTTCTGCACAC,-7429;Transfac,CEBP_C,+,AGTGTGAGGCAAGACCTG,-12861;Jaspar,NF-kappaB,-,GGGAATTTCC,28429;Transfac,EGR3_01,+,CAGCGTGGGAGG,-10034;Transfac,TANTIGEN_B,+,GGGAGGCCGAGGCAGGCAG,-3797;Transfac,SRF_C,-,GCCTTTTTTGGCCCA,12574;Transfac,E4F1_Q6,-,CCTACGTCAC,13357;Jaspar,PPARgamma,-,AGAGGTCAGCGTGACCCCCT,9983;Transfac,HSF_Q6,+,TCCCAGGAGTTTC,-20707;Transfac,EGR1_01,-,TCACGTGGGCGG,6061;Transfac,ETS_Q4,-,TTCCACTTCCTG,32688;Transfac,USF_C,+,CCACGTGA,-6054;Transfac,E2_01,+,GAACCAGAGACGGTGG,-19973;Transfac,AHRHIF_Q6,-,CGCGTGCGG,11119;Transfac,RFX1_02,+,CTGTAGCCTAAGCAACAG,-22798;Transfac,BARBIE_01,-,TTCAAAAGGTGAGGG,28660;Transfac,FXR_IR1_Q6,+,GGATGAATGTCCC,-28051;Transfac,HNF3ALPHA_Q6,-,TGTTTGTTTTG,4737;Transfac,STRA13_01,-,GCCTCACGTGACTC,7198;Transfac,AHR_Q5,+,GTGGCGTGTGC,-21067;Transfac,ZF5_01,-,GGGCGCGG,6795;Jaspar,p65,-,GGGAATTTCC,28429;Transfac,FREAC3_01,-,GGCATGTAAATAAAGA,23069;Transfac,ATATA_B,+,GTATATAAGC,-31222;Transfac,ACAAT_B,+,GATTGGTGG,-26027;Transfac,AP4_Q5,+,CTCAGCTGGC,-13970;Transfac,AP4_Q6,+,CTCAGCTGGC,-13970;Jaspar,Yin-Yang,-,GCCATC,3377;Transfac,ZTA_Q2,-,TCACAGTGACTCA,14023;Transfac,E12_Q6,+,GGCAGGTGCCA,-7403;Transfac,ELK1_02,+,GCTGCCGGAAGGGA,-8752;Transfac,MYC_Q2,+,CACGTGG,-10864;Transfac,LBP1_Q6,-,CAGCTGC,2984;Transfac,TFIII_Q6,+,AGAGGGAGG,-19953;Transfac,LMO2COM_02,+,CAGATAGGG,-43;Transfac,LMO2COM_01,-,CCCCAGGTGTTG,7655;Transfac,SMAD_Q6,-,AGACTCCCC,9856;Transfac,MAF_Q6,+,TGAGGGCAAGTTGGCA,-34778;Jaspar,cEBP,-,TGGCGCAACCTT,38390;Jaspar,c-REL,+,GGGGAATTCC,-23710;Transfac,MUSCLE_INI_B,-,TCCCCCCACCACCCCCTCCCA,30643;Transfac,AP4_Q6_01,+,GCCAGCTGT,-36895;Transfac,DR3_Q4,+,CATCCCCTTCCTGACCCCTCC,-4972;Transfac,STAT5A_04,-,CACTTCCG,16011;Transfac,ATF4_Q2,-,GCTGACGCCACG,4915;Transfac,SPZ1_01,-,GGTGGAGGGATGGGG,16533;Jaspar,TCF11-MafG,+,CATGAC,-3852;Transfac,PAX2_02,+,CACAAACCC,-23836;Transfac,LUN1_01,+,TCCCAGCTACTTGGGAG,-3918;Transfac,PAX2_01,-,CCCTGTCACTCAGGATGGA,20254;Transfac,MAZR_01,-,TGGGGAGGGGCAC,27106;Transfac,MYOGNF1_01,+,AATCCTTTCAGTTTGGGACGGAGTAAGGC,-7790;Transfac,HSF2_01,-,GGAAGCTTCG,13805;Transfac,T3R_01,+,CTGGGAGGTCACGGCT,-21588;Transfac,ZIC3_01,+,TGGGGGGTC,-13048;Transfac,ISRE_01,+,CAGTTTCTCTTCCTG,-29546;Jaspar,Bsap,+,TGGTCAACGCAGCAGAGCGG,-6478;Transfac,CDXA_02,+,ATTACTG,-16382;Transfac,CREB_Q4_01,+,CCGTGACGTAG,-13346;Transfac,ARNT_02,+,CGAGAGTCACGTGAGGCTGA,-7182;Transfac,HOGNESS_B,-,GTGGTGGCTCACGCCTGTAATCCCAGCACT,8124;Transfac,ARNT_01,-,CAGCTCACGTGGGCGG,6065;Transfac,HIF1_Q3,-,GCCCGCGTGCGGCC,11122;Transfac,LFA1_Q6,-,GGGGTCAG,7534;Transfac,GR_Q6,-,GGGCCTCGCTCTGTTGTCC,27466;Transfac,TEF1_Q6,+,GGAATG,-1360;Transfac,BACH1_01,-,GCTATGAGTCACCAC,1540;Transfac,TBP_Q6,+,TTTATAC,-8715;Transfac,E47_02,-,AATTACAGGTGTACGC,21546;Transfac,CP2_02,+,GCTGGGCTGAGCCAC,-6680;Transfac,E47_01,-,AGGGCAGGTGGCTCC,5145;Transfac,MEIS1_01,+,GAGTGACAGGGC,-20244;Transfac,PR_01,-,TGTTGAGGAGAATGCTGTTCTCATTGT,36718;Jaspar,MZF_1-4,+,TGGGGA,-2671;Transfac,OCT1_07,+,TTTATGGTAATT,-31767;Jaspar,Androgen,-,TTTGGCACAGCATGTACCTGTC,34465;Transfac,ZID_01,+,CAGCTCCATCACC,-24971;Jaspar,Pax6,+,TTCACGCTTTAGTT,-2658;Transfac,AREB6_02,+,ACACACCTGTAG,-3906;Transfac,AREB6_03,-,GTGCACCTGTAG,1658;Transfac,PAX_Q6,+,CTGGAAATCAC,-14033;Transfac,RREB1_01,+,CCCCAAAAAACCCT,-1014;Transfac,MEF2_01,-,GGCTAAAACTACCCCT,35670;Transfac,LPOLYA_B,+,CAATAAAG,-22981;Transfac,MEF2_03,-,TAGGTGCCTATAAATAGCATAG,31727;Transfac,ER_Q6,-,AGAGGTCAGCGTGACCCCC,9983;Transfac,MYB_Q6,-,CCCAACTGGC,7236;Transfac,PPARG_02,+,TTCCAGGTGAAGGTGGCCCACTT,-5598;Transfac,HFH4_01,-,TTATGTTTGTTTA,382;Transfac,HEB_Q6,-,GCCAGCTG,13979;Transfac,PPAR_DR1_Q2,+,TGACCTCTGTCCA,-10853;Transfac,OLF1_01,+,CAAGGTTCCCTAGAGAAATGGC,-35076;Transfac,MYOD_01,+,ACACAGGTGGTG,-5933;Transfac,CREBP1_Q2,-,GCTGACGCCACG,4915;Transfac,NERF_Q2,+,TTGCAGGAAGTCAGGGAC,-27797;Transfac,IRF_Q6,+,GTCAGTTTCTCTTCC,-29544;Transfac,XPF1_Q6,+,TCTGGGCAAC,-32109;Transfac,GEN_INI3_B,-,CCTCATTC,17236;Transfac,STAT6_02,+,GCCTTCCT,-7817;Transfac,AR_01,+,GGTACATGCTGTGCC,-34448;Transfac,NFKAPPAB_01,-,GGGAATTTCC,28429;Jaspar,HNF-1,-,GGTTAATAATTATC,35227;Transfac,EGR_Q6,+,GTGGGGGCAAG,-11163;Transfac,LYF1_01,+,TTTGGGAGG,-3584;Transfac,PPARA_01,-,CTGCCCCAGGCCAAATTTCT,12377;Transfac,PPARA_02,-,TGGGGTCAGGCAGGGCTGG,7535;Transfac,COUP_DR1_Q6,+,GGACCTTTGGCTT,-38525;Transfac,GATA1_02,-,TTCTAGATAGGGGC,21667;Transfac,VDR_Q3,-,GAGGGAATGGGGAGA,8449;Transfac,T3R_Q6,+,CCTGTCCTC,-6382;Transfac,VDR_Q6,+,CTGCCTGACCCC,-7523;Transfac,LXR_Q3,-,TGGGGTGACCCTGGTGCG,5511;Jaspar,FREAC-4,+,GTAAACAT,-20345;Transfac,LXR_DR4_Q3,+,TGACCGTCATTAAACC,-8569;Transfac,YY1_02,-,CCTGTGCCATCCAGGCTGGA,14512;Transfac,SP1_01,+,AGGGCGGGGC,-11204;Transfac,AP2_Q6_01,+,CGGCCCCCAGGCC,-4872;Transfac,TCF11_01,-,GTCATTCAGGACC,33780;Transfac,TAL1BETAE47_01,-,GGGGACAGATGGCAGT,25058;Transfac,PAX6_Q2,-,CTGACCTTGAACTC,20070;Transfac,SP3_Q3,-,AGCACTGTGGGAGG,2620;Transfac,SEF1_C,+,GGCCCCCAGGCCTGCGTTC,-4873;Transfac,NFKB_Q6_01,+,GACAAGGAAATTCCCG,-28415;Transfac,ZIC2_01,+,AGGGTGGTC,-27629;Transfac,AREB6_01,-,TACTCACCTGAGT,8388;Transfac,AP2_Q6,+,GGCCCCCAGGCC,-4873;Transfac,HNF4_DR1_Q3,+,TGACCTCTGTCCA,-10853;Transfac,NMYC_01,-,TCCCACGTGGAC,10872;Transfac,AP2_Q3,-,GCCCCCAGCCTTAGGC,22344;Transfac,MYOGENIN_Q6,+,GGCAGCTG,-5067;Transfac,CAP_01,-,TCAGCCCC,36304;Jaspar,c-ETS,+,CTTCCG,-3700;Jaspar,Staf,-,GGTTTCCCAGGGGGCAGTGC,14095;Jaspar,n-MYC,+,CACGTG,-6055;Jaspar,MEF2,+,CTATTTATAG,-31711;Transfac,PAX9_B,-,GTCACCCAGGGTGGAGTGCAGTGA,21178;Transfac,ER_Q6_02,+,GAGGTCACGGC,-21592;Jaspar,HLF,-,GGTTACACAATT,21743;Jaspar,GATA-3,+,AGATAG,-44;Transfac,MZF1_01,+,AGTGGGGA,-6218;Jaspar,Irf-1,-,GATAGTGAAACC,21815;Transfac,E2_Q6,+,GAACCAGAGACGGTGG,-19973;Transfac,SP1_Q6_01,+,AGGGCGGGGC,-11204;Transfac,CREB_Q2,+,CGTGACGTAGGG,-13347;Transfac,CREB_Q3,-,CGTCAG,778;Transfac,NFKB_C,-,AGGGATTTTCCT,20047;Transfac,CREB_Q4,+,CGTGACGTAGGG,-13347;Transfac,SREBP1_01,+,GATCACCTGAG,-4565;Jaspar,Ahr-ARNT,+,CGCGTG,-9987;Jaspar,SRF,-,GCCCATATATGA,37496;Transfac,DR4_Q2,-,CGGCCTCTCCAGACCCA,11714;Transfac,SP1_Q4_01,+,CAAGGGCGGGGCC,-11202;Transfac,TTF1_Q6,+,CCCCCAAGTGTG,-6842;Transfac,ATF_01,+,CCGTGACGTAGGGT,-13346;Transfac,HOXA3_01,+,CCTAATGGG,-35670;Transfac,POU6F1_01,+,GCATAATTTAT,-35917;Transfac,CREB_Q2_01,+,CTTGACGTCAGGAG,-38209;Transfac,GABP_B,-,CCGGGAAGAGCA,19270;Transfac,AHRARNT_01,+,GGAGGGTAGTGTGCCC,-27057;Transfac,DR1_Q3,-,TGGACAGAGGTCA,10865;Transfac,MZF1_02,-,TGGAGAGGGGCAA,19435;Transfac,P300_01,+,TCAAGGAGTGGGTG,-6194;Transfac,DELTAEF1_01,-,ACTCACCTGAG,8387;Jaspar,USF,+,CACGTGG,-10864;Transfac,CMYB_01,+,TACAAAGGCGGTTGGGAG,-11310;Transfac,PADS_C,-,TGTGGTCTC,4001;Jaspar,Chop-cEBP,-,GGGTGCAATGGC,21908;Transfac,DBP_Q6,+,AGCACAC,-6111;Transfac,NFKAPPAB65_01,-,GGGAATTTCC,28429;Transfac,AP2GAMMA_01,-,GCCTGGGGG,4883;Transfac,AHR_01,-,GCCCAGGCTGGAGTGCAA,18623;Transfac,TAL1BETAITF2_01,-,GGGGACAGATGGCAGT,25058;Transfac,PITX2_Q2,+,TGTAATCCCAA,-3780;Transfac,CAAT_C,+,GCCCAATAACCAGCTCCTCGCTGAT,-20432;Transfac,IK2_01,+,CTTTGGGAAGGC,-38457;Transfac,MIF1_01,+,TGGGTGCAGGGCCGCTGG,-7352;Transfac,IK1_01,+,GCTTGGGAAGGCC,-12009;Transfac,NFKB_Q6,+,ATGGGAATCTCCTC,-19067;Jaspar,Tal1beta-E47S,+,GGAACATCTGTT,-35130;Transfac,VJUN_01,+,GTGATGATGTCATTGC,-6140;Transfac,PAX5_02,+,GGAGTGCAATGTGAGCCGAGACCACACA,-3976;Transfac,PAX5_01,-,TCTTGGCTCACTGTAGTGTAGACTTCCC,18984;Transfac,BRACH_01,-,AGAATCACATGTAGGTGCCACAGT,16237;Transfac,CETS1P54_02,-,CCACCGGATGTGG,5441;Transfac,MAF_Q6_01,-,GGCTGAGTCAA,24942;Transfac,TAXCREB_02,+,GTGACCCACACCCTA,-28621;Jaspar,Pax-2,-,CGTCACGG,13353;Transfac,COMP1_01,+,TGTTATCAATGACAATGCGCGCCC,-28488;Transfac,CREL_01,+,GGGGAATTCC,-23710;Transfac,SP1_Q2_01,-,CCCCACCCCC,8399;Jaspar,c-MYB_1,+,GGCCGTTG,-11773;Transfac,SMAD3_Q6,-,TGTCTGTCT,16822;Transfac,E2A_Q6,+,CACCTGCC,-5136;Transfac,MYCMAX_03,+,CGAGAGTCACGTGAGGCTGA,-7182;Transfac,CHCH_01,+,CGGGGG,-6696;Transfac,E2A_Q2,-,GCACCTGCCTCAGT,7411;Transfac,BEL1_B,-,AAAGTGCTGAGATTACAGGCATAAGCCA,17103;Transfac,NRSE_B,+,CTCAGCACCTTGGCCAGCTCC,-24957;Transfac,MAZ_Q6,-,GGGGAGGG,16549;Transfac,ZIC1_01,+,TGGGGGGTC,-13048;Jaspar,RORalfa-1,+,TTCAAGGTCA,-20060;Transfac,NF1_Q6,+,TGCTGGCAGGCAGGCAGA,-12343;Transfac,MINI20_B,+,ACCTCCCACCATGGAGGAGGA,-5205;Transfac,VMW65_Q6,+,TCTCATTA,-25555;Transfac,NFKAPPAB50_01,+,GGGGAGTCCC,-5241;Jaspar,RREB-1,-,CCCCCCACCACCCCCTCCCA,30642;Jaspar,NRF-2,+,GCCGGAAGGG,-8755;Transfac,RFX1_01,+,TAGGCACCTAGTAACAG,-31718;Transfac,GNCF_01,+,CAGGAGTTCAAGGTCAGC,-20054;Jaspar,RXR-VDR,-,GGGTCACAGAGATCA,28627;Transfac,NRSF_01,+,CTCAGCACCTTGGCCAGCTCC,-24957;Transfac,USF_Q6_01,+,GCCCACGTGAGC,-6052;Transfac,P53_01,+,GGACATGGTGGCACATGTCT,-22689;Transfac,WHN_B,+,AGGGACGCCTT,-6534;Transfac,MINI19_B,-,GCAAGGAGCCACACAGCAGGA,13854;Transfac,GKLF_01,+,AAAGGAAGGAAGGG,-35999;Transfac,HNF4_01_B,+,GGGGGCAAAGGTAGG,-22339;Transfac,YY1_Q6,-,GCCATCTTG,18004;Jaspar,p53,-,CAGGACAAGTTCGAGCATCT,2978;Jaspar,p50,-,GGGGGTTCCCG,15798;Transfac,GATA2_02,-,GGAGATAAGA,33994;Transfac,GRE_C,+,GTCACACCCTGTCCTC,-6375;Transfac,FXR_Q3,+,CAAGGGCAGCAACC,-13934;Transfac,MYCMAX_B,-,GCCATGTGCC,30955;Transfac,NFE2_01,-,AGCTGAGGCAC,13976;Transfac,CACBINDINGPROTEIN_Q6,+,GGGGGTGGG,-8390;Transfac,MYOD_Q6,+,TGCACCTGTC,-6277;Transfac,STAF_02,+,ACATACCATCATGCCTGGCTA,-24189;Transfac,STAF_01,-,AGTTCCCGTAGTGCCTGACGGT,5931;Transfac,GATA3_02,-,GGAGATAAGA,33994;Jaspar,Myf,+,AGGCAGCAGGAG,-8418;Transfac,NRF2_01,+,GCCGGAAGGG,-8755;Transfac,GATA1_01,+,GGGGATGGGG,-6520;Transfac,ICSBP_Q6,+,GAAGAGAAACTG,-6711;Transfac,CETS1P54_01,-,ACCGGATGTG,5439;Transfac,TCF11MAFG_01,+,CTGTTGTGAGGCAGCAGTTGTG,-12574;Transfac,CACCCBINDINGFACTOR_Q6,+,AATCAGCTGGGTGTGG,-18121;Transfac,SMAD_Q6_01,+,TAGTCAGACAG,-34438;Transfac,GC_01,+,CAAGGGCGGGGCCT,-11202;Transfac,FOXM1_01,+,AGATGGAGT,-3171;Transfac,ARP1_01,-,TGAACTCCTGACCTCT,3835;Transfac,NGFIC_01,-,TCACGTGGGCGG,6061;Jaspar,Gklf,+,AAAGGGAAGG,-35981;Transfac,ERR1_Q2,+,AGTTCAAGGTCAGC,-20058;Jaspar,MZF_5-13,-,GGAGGGGGAG,8091</motif>
</e>
</a>
Other examples could be found in the previous version.